PrimerChecker
A PCR primer validation tool that combines three-source genomic search with local thermodynamic analysis.
Problem
Primer validation is a chore. You need to check thermodynamic properties (Tm, GC content, hairpins, dimers) and specificity across the genome, which today means bouncing between a local tool for the maths and at least two websites (UCSC BLAT, NCBI BLAST) for the searches. Each step is cheap; the context-switching isn't.
Approach
A Next.js frontend on a FastAPI Python backend. Local analysis modules compute melting temperature via nearest-neighbour thermodynamics (SantaLucia, 1998), plus GC content, hairpin risk, and self- and cross-dimer checks, with a traffic-light verdict (green / amber / red) rolled up at the top of the report. Genomic specificity fans out in parallel to three sources — NCBI Primer-BLAST, NCBI BLAST E-utilities, and EBI BLAST + Ensembl REST — and a confidence score (high / medium / low) is assigned based on how many sources agree. Recent searches are cached in localStorage (max 8, deduped by primer pair and mode) so prior queries are one click away.
Stack
Synopsis
The scope is primer validation end-to-end — local thermodynamic analysis (Tm, GC, hairpin, dimer), cross-database specificity against three genomic sources, and a unified traffic-light verdict. The purpose is to collapse the pre-order validation chore from a tour of separate websites and manual cross-referencing into a single panel a scientist can read in under a minute.
Gallery
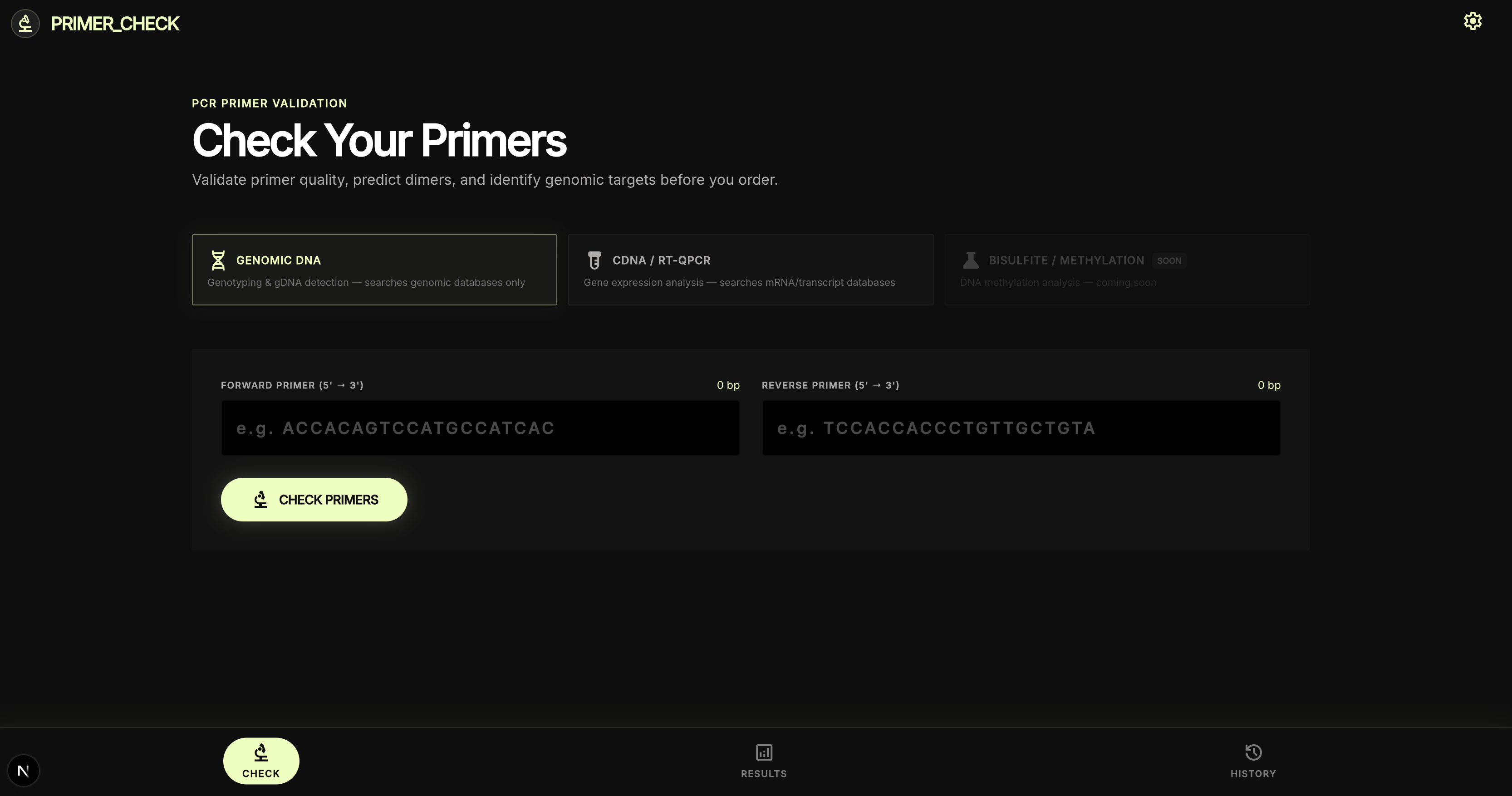
Primer input and target selection
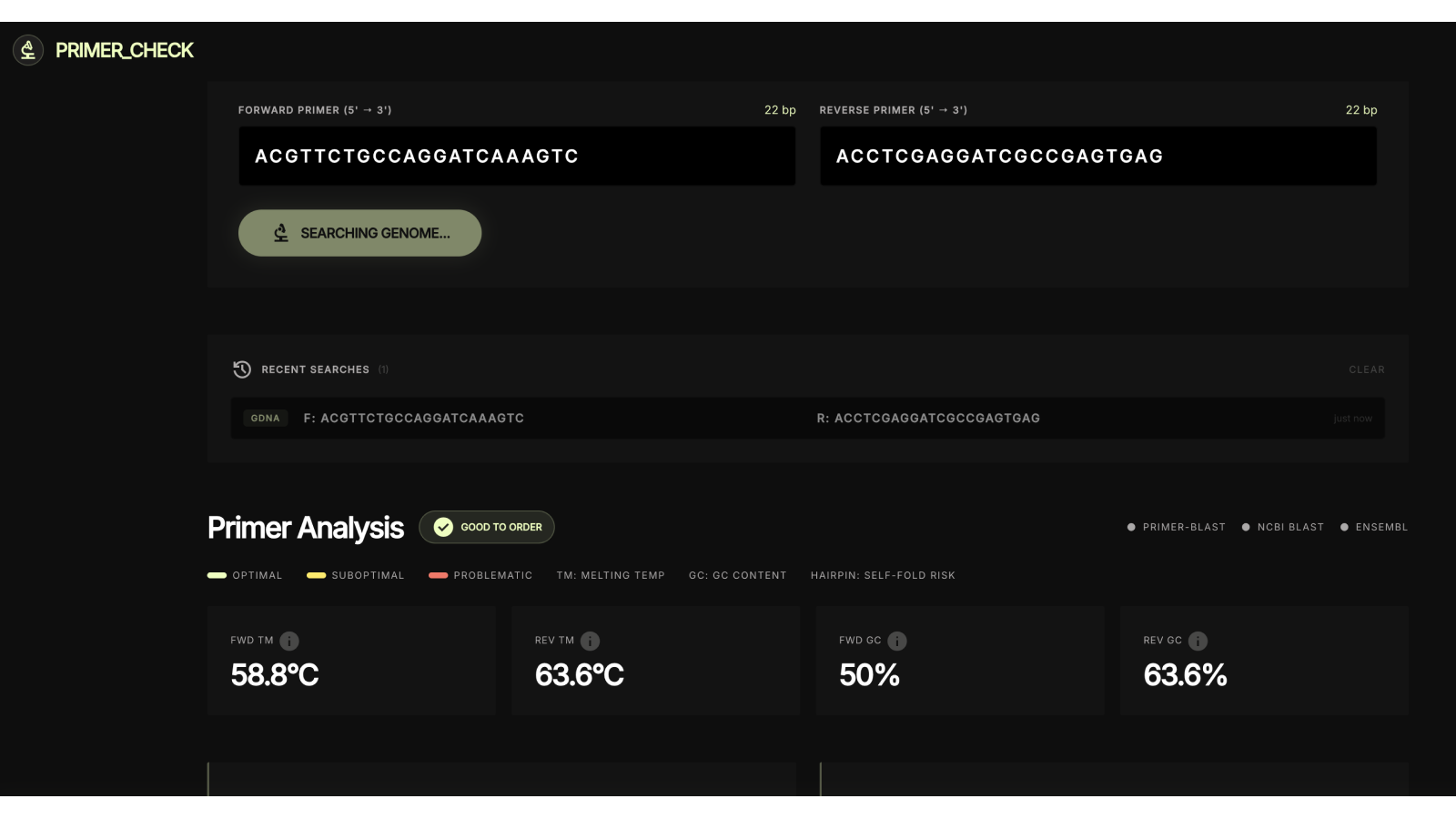
Genome search kicks off in parallel
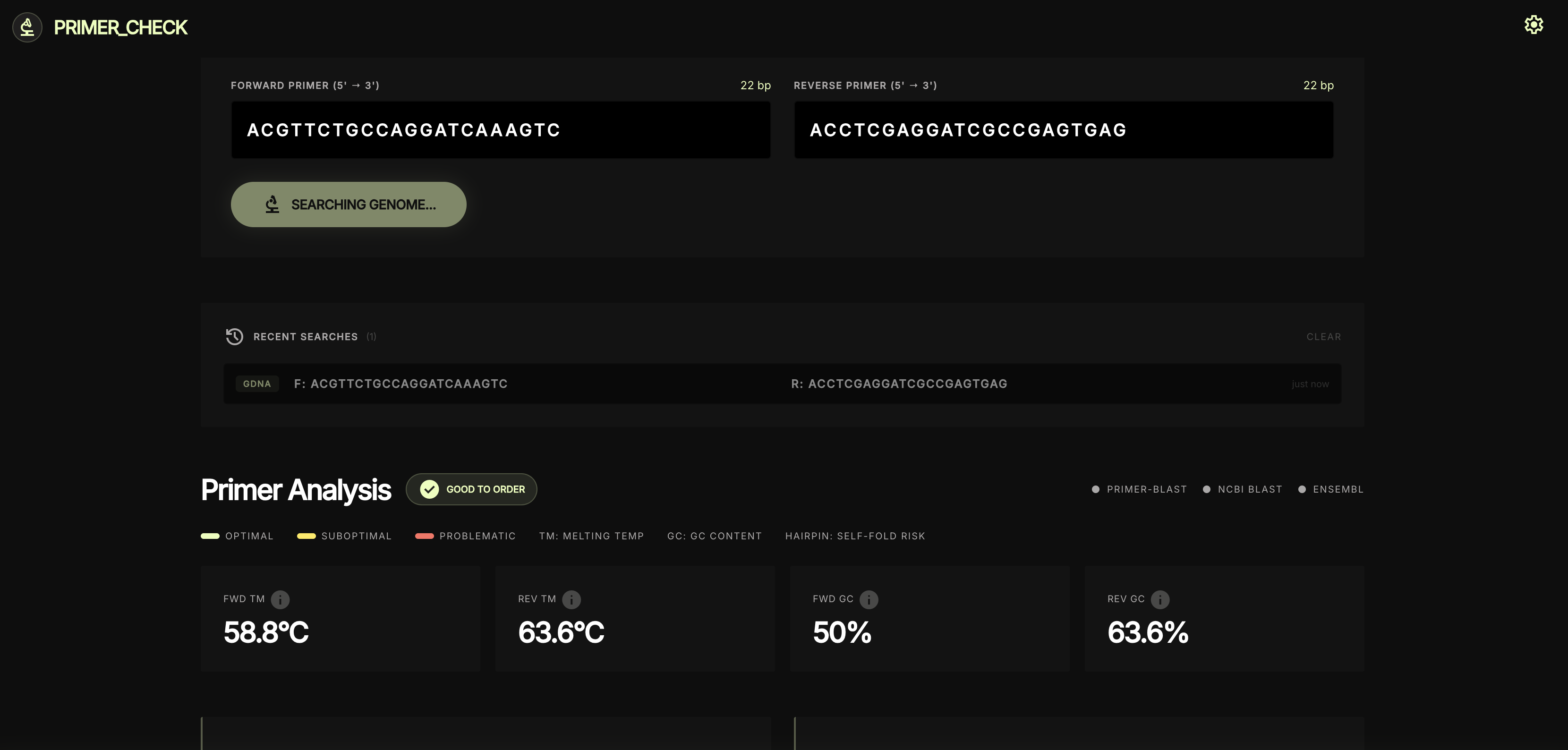
Per-primer thermodynamics
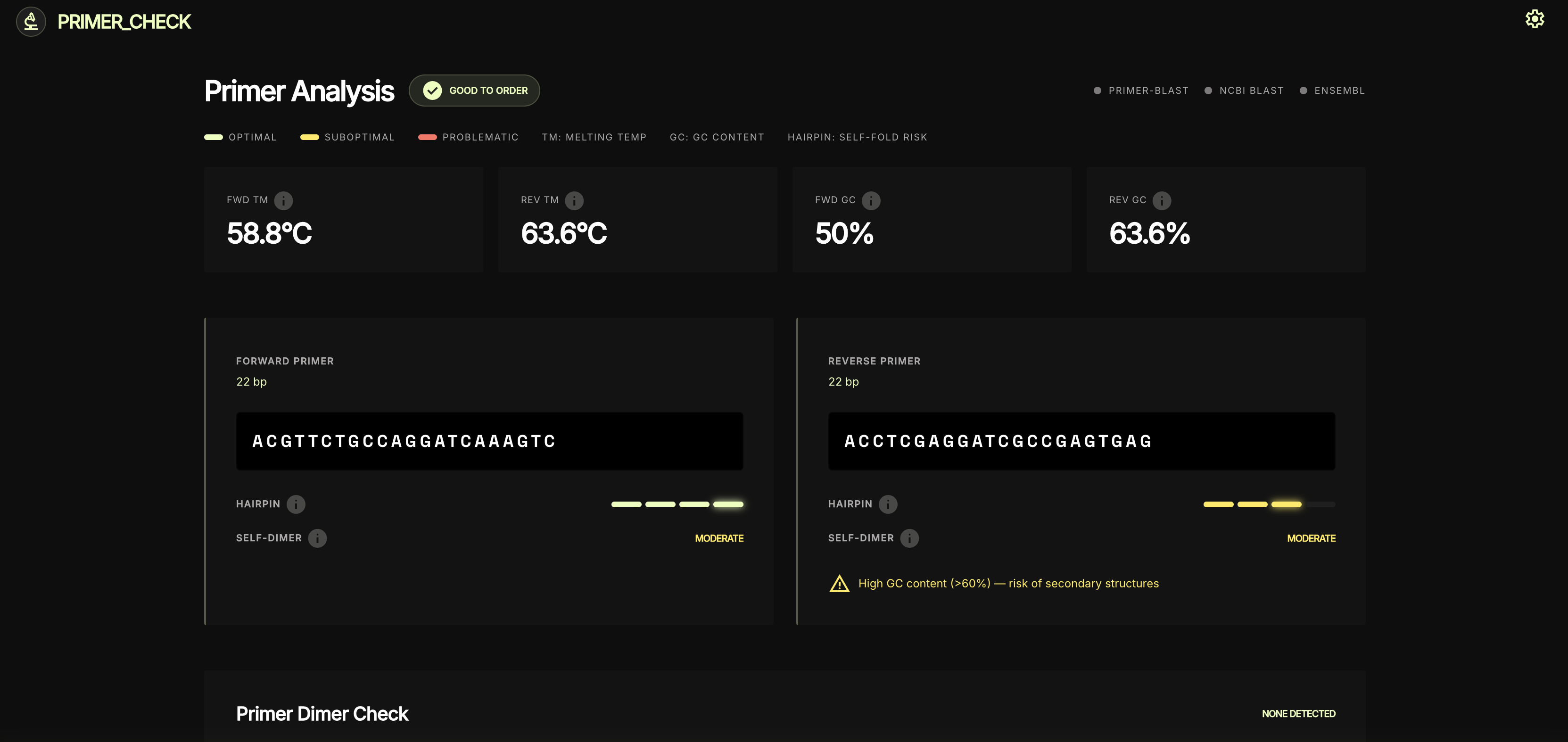
Dimer check + specificity status